Table of Contents
Definition / general | Microscopic (histologic) description | Positive stains | Negative stains | Molecular / cytogenetics description | Molecular / cytogenetics imagesCite this page: Mihova D. t(12;21)(p13;q22); ETV6::RUNX1. PathologyOutlines.com website. https://www.pathologyoutlines.com/topic/leukemiapreBALLt1221.html. Accessed April 23rd, 2024.
Definition / general
- 20 - 30% of childhood preB ALL; most common translocation (Chin Med J (Engl) 2003;116:1298) but not infants; also 3% of adults
- 5 different patterns of gene expression involving 14 genes, can detect with gene chip (BMC Genomics 2007;8:385)
- Excellent prognosis due to good response to chemotherapy; 90% remissions; relapses occur later than other ALL
- Persistence of TEL-AML1 transcripts is not necessarily related to relapse (Pediatr Int 2003;45:275)
Microscopic (histologic) description
- No distinct morphology
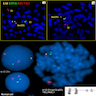
Molecular / cytogenetics description
- Fusion of TEL/ETV6 and AML1/RUNX1/CBFA2 genes
- Detect with FISH or PCR; not found by conventional cytogenetics (i.e. are cryptic) because rearranged segments are too small
- May have no other molecular abnormalities, almost never > 50 chromosomes