Table of Contents
Definition / general | Essential features | ICD coding | Epidemiology | Pathophysiology | Clinical features | Laboratory | Prognosis | Case reports | Treatment | Microscopic (histologic) description | Microscopic (histologic) images | Peripheral smear images | Positive stains | Negative stains | Molecular / cytogenetics description | Differential diagnosis | Board review style question #1 | Board review style answer #1 | Board review style question #2 | Board review style answer #2Cite this page: Kaseb H, Hudnall SD. BCR::ABL1-like. PathologyOutlines.com website. https://www.pathologyoutlines.com/topic/leukemiaprebbcrabl1like.html. Accessed April 19th, 2024.
Definition / general
- Introduced as a provisional entity in the WHO 2017 (Swerdlow: WHO Classification of Tumours of Haematopoietic and Lymphoid Tissues, 4th Edition, 2017)
- Does not have BCR-ABL1 translocation but shows a pattern of gene expression similar to that seen in ALL with BCR-ABL1
Essential features
- Exhibits National Cancer Institute (NCI) high risk features: (a) WBC ≥ 50,000/microL or age ≥ 10 years; up to 13 years if treated on a COG protocol; (b) persistent, postinduction, minimal residual disease; and (c) genetic alterations in kinase signaling including cytokine receptors and tyrosine kinases (Pediatr Blood Cancer 2017;64:10.1002)
- Important to recognize because:
- Associated with a poor prognosis
- Some are sensitive to tyrosine kinase inhibitors similar to BCR-ABL1 (Eur J Cancer 2017;82:203, Cancer Cell 2016;29:186)
- Use of tyrosine kinase inhibitors with treatment protocols may improve outcome (Eur J Cancer 2017;82:203)
- Identified in 15 - 20% of patients with B ALL (Lancet Oncol 2009;10:125, N Engl J Med 2009;360:470)
ICD coding
- ICD-10: C91.00 - acute lymphoblastic leukemia not having achieved remission
Epidemiology
- Relatively common subtype, representing 7 - 25% of B ALL patients (Swerdlow: WHO Classification of Tumours of Haematopoietic and Lymphoid Tissues, 4th Edition, 2017)
- Reported in both pediatric and adult patients
- More common in Down syndrome patients; uniquely shows CRLF2 translocation
Pathophysiology
- Harbors a large number of kinase activating gene rearrangements primarily involving the ABL class, JAK / STAT or RAS pathway associated signaling pathways
- Common genes involved include ABL1, ABL2, CRLF2, CSF1R, EPOR, NTRK3, PDGFRB, JAK1, JAK2, JAK3, FLT3, IL7R, SH2B3 and IKZF1
- These genes encode proteins involved in B cell development, proliferation and differentiation, cell cycle regulation and cell signaling
- Overexpression of cytokine receptors such as cytokine receptor-like factor 2 (CRLF2) occurs in both ALL with BCR-ABL1-like / Philadelphia-like and other B ALL categories (Eur J Cancer 2017;82:203)
Clinical features
- B ALL in general usually presents with symptoms related to bone marrow suppression by lymphoblasts
- Patients can present with anemia, leucopenia or thrombocytopenia or a combination of these
- Symptoms include bruising or bleeding due to thrombocytopenia, pallor or fatigue due to anemia and recurrent infections caused by neutropenia / leucopenia or bone pains
- May also present with lymphadenopathy (> 10 mm in single dimension of the lymph node), hepatomegaly or splenomegaly
- B ALL with BCR-ABL1-like and other B ALL with recurrent genetic abnormalities show no unique clinical presentation or microscopic findings
- Further, similar to some other B ALL with recurrent genetic abnormalities, B ALL with BCR-ABL1-like shows no unique immunophenotypic profile (Pediatr Blood Cancer 2017;64:10.1002)
- Essentially a diagnosis of exclusion
- Identifying patients with B ALL with BCR-ABL1-like can influence the management of the patient
- Those with PDGFRB translocation can benefit from tyrosine kinase inhibitors, while patients with JAK translocations may benefit from JAK inhibitors (Pediatr Blood Cancer 2017;64:10.1002)
- Adults tend to have poor outcome even with high intensity chemotherapy regimens
Laboratory
- B ALL with BCR-ABL1-like patients do not show a BCR-ABL1 fusion protein expressed from t(9;22)(q34;q11.2); however, they have a gene expression profile similar to ALL with BCR-ABL1
- Almost none of the genetic alterations are detected by standard genetic diagnostic methods such as conventional karyotyping and FISH because these genetic alterations are diverse and often cryptic
- To date, there are over 60 different identified rearrangements and approximately 16 different targetable genes
- Patients can only be diagnosed by gene expression panels (genome wide Affymetrix gene expression arrays):
- Currently 2 gene expression signatures have been verified for diagnosis of B ALL with BCR-ABL1-like / Ph-like
- BCR-ABL1-like gene expression array consists of 110 expression probe sets and was originally developed to classify B ALL in a population based Dutch / German cohort
- Gene expression of a group of cases were similar to ALL with BCR-ABL1 and therefore named the category ALL with BCR-ABL1-like
- These had an overall unfavorable outcome (Lancet Oncol 2009;10:125)
- Philadelphia-like signature was first demonstrated in cases of B ALL with IKZF1 deletions in a high risk United States cohort (N Engl J Med 2009;360:470)
- Differences between these 2 gene expression signatures can be explained by their different origins and discovery cohorts
- Few reference laboratories are currently offering low density microarray classifiers (8 or 15 gene) that may help screen patients
- One of these reference laboratories is Tricore reference laboratories (BCR-ABL1 like ALL (Ph-like ALL): TriCore Reference Laboratories [Accessed 8 May 2019])
- Some institutions and reference laboratories have developed multiplex FISH testing that may help screen cases
- Some institutions such as St. Jude utilize transcriptome and whole exome sequencing
- Flowcytometry:
- There is evidence that B-ALL with BCR-ABL1-like present as common-ALL on IPT, however this maturation stage is not distinct of the entity and is common in other B-ALL subt
- ypes B-ALL with BCR-ABL1-like patients with CRLF translocations show high levels of surface expression of the protein on flow cytometry, which can be used as a screening tool
Prognosis
- Prognostic factors include age, white blood cell count, immunophenotype, genetics and detection of measurable residual disease
- Numerous stratification schemes to assess risk in B ALL
- NCI risk group:
- NCI standard risk group: WBC < 50,000/microL and age 1 to < 10 years
- NCI high risk group: WBC ≥ 50,000/microL or age ≥ 10 years (up to 13 years if treated on a COG protocol)
- One scheme stratifies patients into standard risk and high risk based on age (< 10 years = standard, > 10 years = high) and WBC count (< 50,000/mm3 = standard, > 50,000/mm3 = high)
- Recurrent genetic abnormalities are identified in the majority of B ALL cases
- Prognosis of B ALL with BCR-ABL1-like is generally unfavorable
Case reports
- 10 year old boy treated with tyrosine kinase inhibitor for EBF1-PDGFRB fusion (J Clin Oncol 2013;31:e413)
- 16 year old boy treated with tyrosine kinase inhibitor for EBF1-PDGFRB fusion (Haematologica 2013;98:e146)
- 16 year old girl and 29 year old man with B ALL and novel ABL1 fusion genes (Br J Haematol 2011;153:43)
- 17 year old girl with novel JAK2 F694L mutation treated with JAK2 inhibitor and stem cell transplant (Pediatr Blood Cancer 2017;64:10.1002)
- 20 year old patient with a relapse of Philadelphia chromosome negative ALL who responded to tyrosine kinase inhibitor therapy (Oncol Res Treat 2018;41:550)
Treatment
- B ALL with BCR-ABL1-like is managed by standard combination chemotherapy
- Effective treatment modality for B ALL since the 1950s
- Usually administered in 3 distinct phases (induction, consolidation and maintenance) and should include intrathecal treatment, which is directed to the central nervous system
- Addition of tyrosine kinase inhibitors in B ALL with BCR-ABL1-like depends on the specific genetic alteration and has been shown to improve patient's outcome
- Both imatinib and dasatinib specifically inhibit ABL1, ABL2, PDGFRB and CSF1R kinases; ruxolitinib inhibits JAK / CRLF2 / EPOR alterations (Eur J Cancer 2017;82:203)
- Sensitivity to ruxolitinib in patients with JAK class aberrations is promising and has been successful in 2 JAK class fusion positive patients and 1 JAK2 mutated ALL (Eur J Cancer 2017;82:203)
- Adults tend to have poor outcome even with high intensity chemotherapy regimens; therefore, bone marrow transplantation becomes an important treatment option
Microscopic (histologic) description
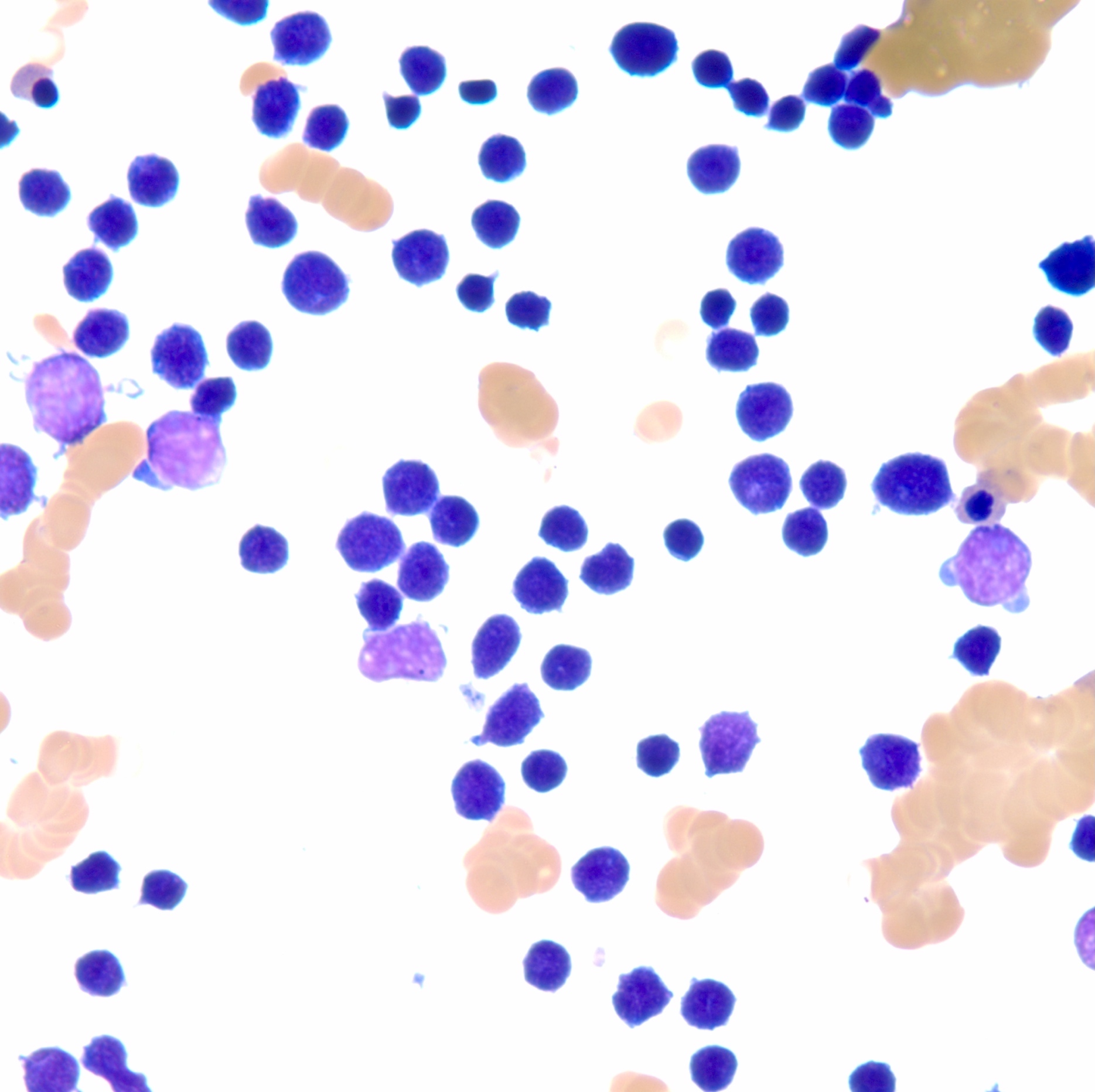
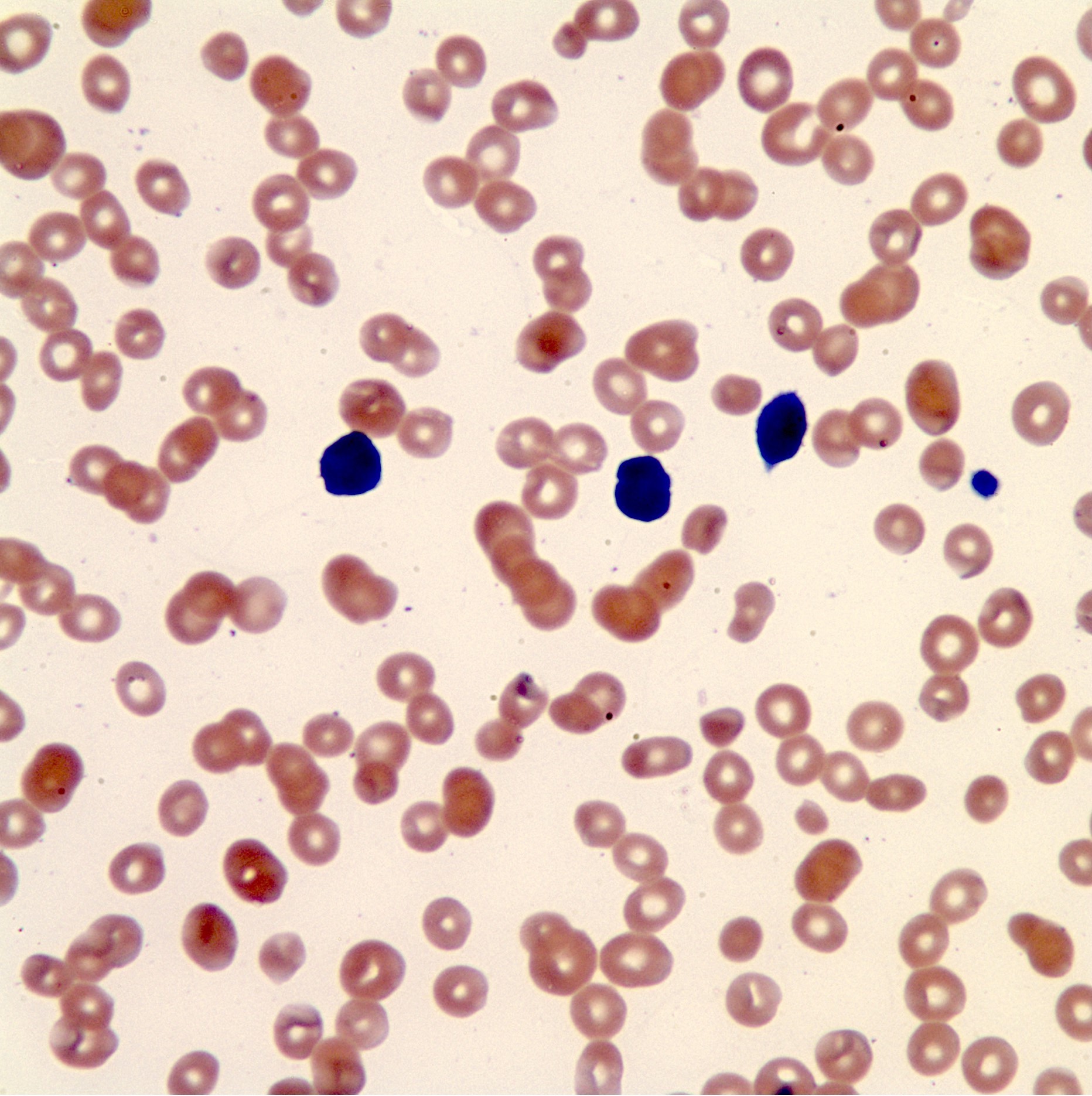
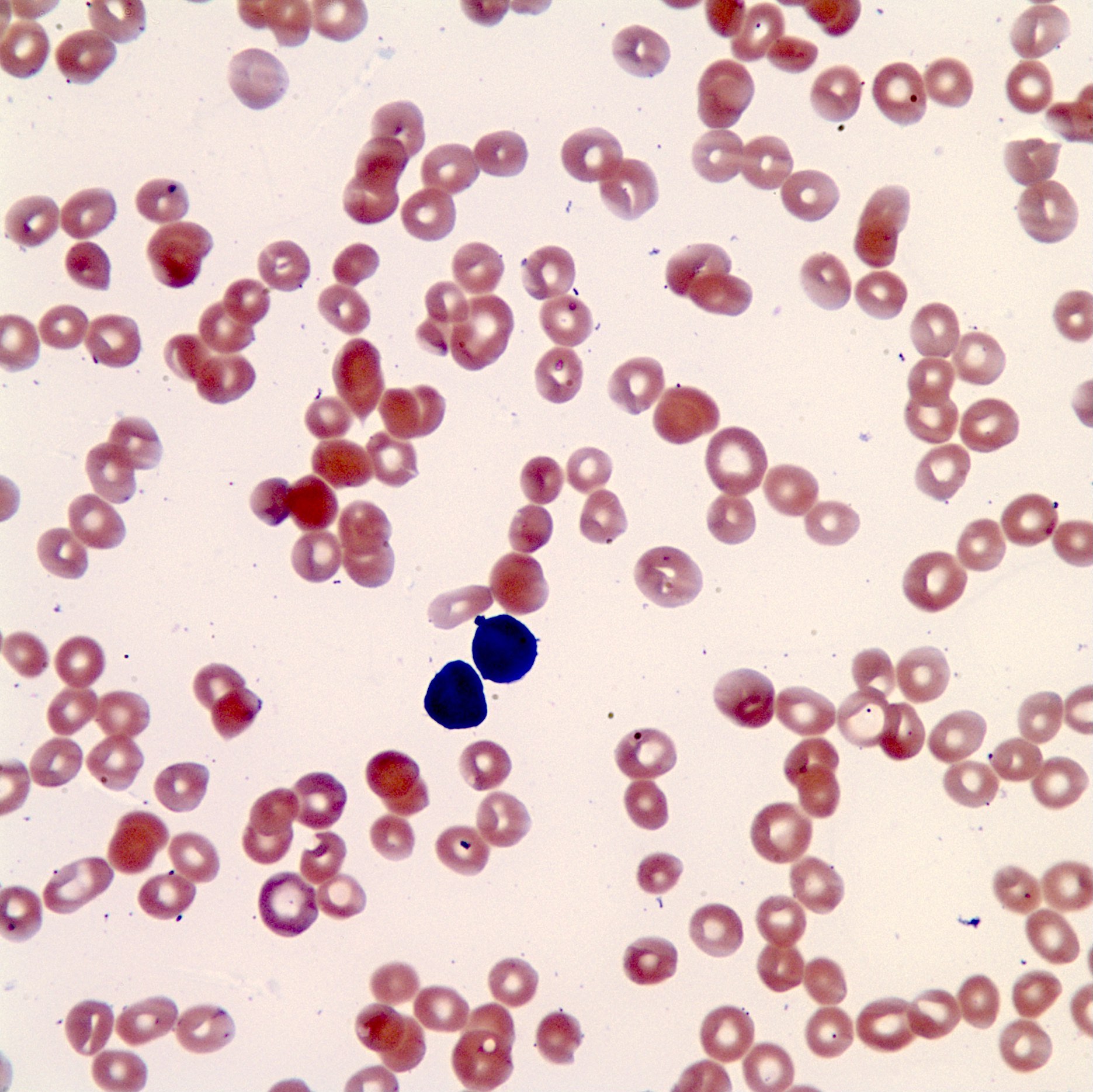
- Blasts have scant agranular cytoplasm, coarse to fine chromatin, often with indistinct nucleoli
- No Auer rods and no dysplastic myeloid cells
Positive stains
Molecular / cytogenetics description
- Gene expression profiling is the gold standard for diagnosis of B ALL with BCR-ABL1-like / Ph-like (Lancet Oncol 2009;10:125, N Engl J Med 2009;360:470)
- Genetic alterations in B ALL with BCR-ABL1-like are harder to detect by standard genetic diagnostic approaches (Eur J Cancer 2017;82:203)
- Common gene rearrangements include CRLF2, EPOR and IGH
- Down syndrome patients uniquely show CRLF2 translocation
- ABL1 oncogene translocation with ABL2, PDGFRB, NTRK3, TYK2, CSF1R and JAK2 have all been reported
- Many cases of B ALL with BCR-ABL1-like may additionally show other deletions or mutations that have a clear role in leukemogenesis such as IKZFA and CDKN2A / B
- FISH and karyotyping are helpful in ruling out other B ALL entities, especially ALL with recurrent genetic abnormalities
- Multiplex FISH testing can be useful in ruling out B ALL with recurrent genetic abnormalities
- Identifying patients with B ALL with BCR-ABL1-like can influence patients' prognosis and management of the patient
- PDGFRB translocation: may benefit from tyrosine kinase inhibitors
- JAK translocations: may benefit from JAK inhibitors
Differential diagnosis
- B lymphoblastic leukemia / lymphoma, NOS: diagnosis should only be rendered after the exclusion of all other entities
- Burkitt lymphoma / leukemia
- B lymphoblastic leukemia / lymphoma with recurrent genetic abnormalities: must perform genetic testing including karyotyping, FISH or gene expression array; the entities in this group include:
- B ALL / LBL with t(9;22)(q34;q11.2); BCR-ABL1
- B ALL / LBL with t(v;11q23.3); KMT2A rearrangement
- B lymphoblastic leukemia / lymphoma with t(12;21)(p13.2;q22.1); ETV6-RUNX1
- B lymphoblastic leukemia / lymphoma with hyperdiploidy
- B lymphoblastic leukemia / lymphoma with hypodiploidy
- B lymphoblastic leukemia / lymphoma with t(5;14)(q31.1;q32.3); IL3-IGH
- B lymphoblastic leukemia / lymphoma with t(1;19)(q23;p13.3); TCF3-PBX1
Board review style question #1
What is the most sensitive test in diagnosing B lymphoblastic leukemia / lymphoma with BCR-ABL1-like / Ph-like?
- Flow cytometry
- Gene expression profiling
- Karyotyping
- Multiplex FISH
Board review style answer #1
B. Only gene expression profiling can diagnose the entity B ALL with BCR-ABL1-like / Ph-like
Comment Here
Reference: BCR-ABL1-like
Comment Here
Reference: BCR-ABL1-like
Board review style question #2
Which of the following is a characteristic of B lymphoblastic leukemia / lymphoma with BCR-ABL1-like / Ph-like?
- NCI high risk features
- NCI intermediate risk features
- NCI low risk features
- NCI risk features is not applicable
Board review style answer #2
A. ALL with BCR-ABL1-like shows the following characteristics: NCI high risk features, persistent postinduction minimal residual disease (MRD), genetic alterations that activate kinase signaling (Pediatr Blood Cancer 2017;64:10.1002). NCI risk stratifications include NCI standard risk group: WBC < 50,000/microL and age 1 to < 10 years; NCI high risk group: WBC ≥ 50,000/microL or age ≥ 10 years (up to 13 years if treated on a COG protocol).
Comment Here
Reference: BCR-ABL1-like
Comment Here
Reference: BCR-ABL1-like