Table of Contents
Definition / general | History | Procedure for DNA purification | Procedure for RNA purification | Tissue preparation | Anion exchange chromatography | Cesium chloride (CsCl) density gradient centrifugation | Ethanol precipitation / salting out | Organic extraction | Silica adsorption | PCR Inhibitors | Commercial DNA extraction machines | Analyzing purity | Diagrams / tables | Videos | Additional referencesCite this page: Shackelford RE. DNA & RNA purification. PathologyOutlines.com website. https://www.pathologyoutlines.com/topic/moleculardnapurintro.html. Accessed April 19th, 2024.
Definition / general
- Extremely pure nucleic acid samples (RNA or DNA) are required by molecular, research and forensic molecular testing techniques such as PCR amplification of target nucleic sequences, sequencing, blotting / strip assays and cloning procedures
- Often the target nucleic acids must initially be isolated from crude samples that are often contaminated with organic or inorganic substances that can inhibit the PCR reaction or other aspects of DNA amplification
- Additionally, many target nucleic acid in crude samples are unstable and easily degraded or chemically modified by DNases, RNases, formaldehyde
- Rapid, efficient sample processing is often vital to insure sufficiently intact nucleic acid for testing protocols
- Contaminating exogenous nucleic acids can enter samples during the isolation procedure, leading to false positive results and possible treatment errors; for these reasons, prompt, effective and cost efficient nucleic acid isolation techniques are vital
History
- Nucleic acids first isolated in 1869 by Friedrich Miescher, who identified a chemical in the nuclei of leukocytes that was rich in phosphate and nitrogen and lacking sulfur; named "nuclein," due to derivation from nucleus
- Later, Albrect Kossel isolated the nonprotein content of nuclein, finding adenine, cytosine, guanine, thymine and uracil from nuclein, giving some information about the molecular structure of DNA
- Other researchers isolated and identified the chemically linked base, sugar, phosphate nucleotide unit and identified sugar and base differences between RNA and DNA
- Initial nucleic acid preparations were crude and mainly depended on various buffered salt solutions with acid and alkali washes to precipitate and isolate nucleic acids
- "Warm alcohol" was used to remove lipids, pepsin was used to digest proteins
- Since these early efforts, enormous progress has been made in nucleic acid isolation and many highly effective techniques are now used for different applications
Procedure for DNA purification
- Initial nucleic acid isolation requires cells lysis or solubilization of nucleic acids in sample
- Nucleic acids are most commonly isolated from whole blood, plasma, white cells, tissue (often formalin fixed or quick frozen), feces, urine
- Break / disperse sample into solution; often involves cell lysis to expose the nucleic acids
- Remove lipid / membrane with various detergents or less commonly alcohols; most samples containing cells will require the separation of cell membranes from nucleic acids
- Remove proteins by digestion with Protinase K or other protease K
- To obtain pure DNA:
- Incubate sample with RNase (to digest / remove RNA); RNA can also be removed by increasing alkalinity, causing internal phosphoester transfer reactions, leading to RNA strand cleavage
- Add divalent cation chelators to bind and sequester Mg2+ and Ca2+, which are often required for DNase activities
- Additional steps depend on sample type; for example, resins in plants samples can make nucleic acid isolation complex and difficult
- Are many specific different nucleic acid isolation protocols; often two or more are combined for better nucleic acid purification, such as organic extraction followed by cesium chloride gradient centrifugation
Procedure for RNA purification
- RNA is less stable than DNA, making its purification more difficult
- 4M guanidinium thiocynate or phenol with SDS are often used to denature RNases naturally present in blood, plasma, body fluids, bacteria, fungi, plants
- Must avoid sample contamination with RNases, which are extremely ubiquitous in environment
- Must use extremely clean reagents, such as diethylpyrocarbonate treated water
- Must keep samples cold to lower RNase activity; isolate pure RNA quickly and efficiently to minimize RNase exposure and avoid a basic pH in all extraction and storage solutions, as RNA rapidly degrades under basic conditions
Poly (A)+ RNA purification
- mRNA represents only 1 - 2% of total cellular RNA but is often analyzed for research and molecular diagnostic purposes
- Most eukaryotic mRNAs carry poly (A)+ 3' terminal tails, which can be used to isolate mRNA over an oligo (poly dT) attached to a solid matrix
- Under the right buffer conditions, cellular mRNA poly (A)+ tails will form RNA-DNA hybrids; the solid phase can then be washed to remove unwanted cellular continents and a later denaturing wash can be employed to remove pure mRNA
Tissue preparation
- Different sources of nucleic acids often require different purification steps, depending on the nonnucleic acid constituents and the amount and type of nucleic acid degrading enzymes present in the sample
- Although not commonly performed, isolation of nucleic acids form plants sources can be very difficult due to contaminating resins
- Two commonly encountered problems are nucleic acid isolation from formalin fixed or formalin embedded tissues and bone samples subjected to decalcification (decal)
- Formalin fixed or formalin embedded tissues:
- Nucleic acids are usually fragmented and cross linked to other biomolecules, making amplification (especially of longer DNA sequences) difficult
- Contains polymerase chain reaction (PCR) inhibitors which must be thoroughly removed prior to amplification
- May have formalin fixation introduced DNA sequence alterations
- Tumor tissue that may require analysis often contains intermixed areas of benign tissue or benign stromal elements, often necessitating histological examination and microdissection to increase the portion of tumor within the sample
- Decalcified bone:
- Nucleic acid isolation from decalcified bone is possible but the nucleic acids are often so degraded by decalcification that the yield is too low and the nucleic acids are too fragmented for successful PCR amplification
- For this reason, many labs do not attempt nucleic acid isolation from decalcified bone
Anion exchange chromatography
- Subtype of ion exchange chromatography (Wikipedia: Ion Chromatography [Accessed 29 May 2018], Tosoh Bioscience: Ion Exchange [Accessed 29 May 2018])
- Rapid and efficient method of nucleic acid purification which removes lipids, proteins and other cellular constituents in one rapid step
- Uses a positively charged solid resin column that captures negatively charged nucleic acids
- Prior to use, the column is equilibrated with a buffer containing anions and the crude nucleic acid preparation (usually a cell lysate) is added to the column
- Negatively charged nucleic acids are preferentially retained in the column while other cellular constituents are washed out, allowing one step purification
- After several washes to remove unwanted cellular constituents, the nucleic acids are removed with a high salt, low pH wash
- Commonly, the elution profiles of RNA and DNA overlap and other steps must be undertaken if pure RNA or DNA is required
- Diethylaminoethyl (DEAE) covalently linked to sepharose (a polysaccharide polymer) is used in anion exchange chromatography
Cesium chloride (CsCl) density gradient centrifugation
- Mixes nucleic acid, CsCl and ethidium bromide and subjects the mix to high speed centrifugation
- Can use for specific bands of different nucleic acid types, which can be removed and further purified with salting out to remove residual CsCl
- CsCl is highly soluble in water and has been used to isolate many different nucleic acid types, including chromosomal, plasmid and organelle (mitochondrial or plasmid) DNAs and different RNA types (rRNA, tRNA or mRNAs)
- Separation is based on nucleic acid weight and is so exact that nucleic acids of the same size and sequence can be separated based on different isotopic labels (example: N14 vs N15)
- Technique has been used since the 1950s and is sensitive enough to separate similarly sized DNA fragments based on differing A-T or C-G content
- Typically, intact cells are collected by low speed centrifugation, lysed in alkaline conditions with a detergent, protease and RNase to solublize lipids and digest proteins and RNA
- Alternatively, RNA can be harvested by this method, either by isolating the specific RNA nucleic acid band or by predigesting the DNA in the sample with DNase
- Sample is often partially purified by short, low velocity centrifugation to remove flocculent materials
- Supernatant is then loaded over a CsCl buffer solution and centrifuged at ultra high speeds, causing the CsCl to form a gradient into which the nucleic acids migrate until they reach a point of neutral buoyancy (the isopycnic point)
- CSCl centrifugation results in extremely pure nucleic acids
- Following ultracentrifugation, rotor is stopped slowly with brakes off to minimize possible disturbances to nucleic acid bands
- Ethidium bromide is very hydrophobic and is removed from DNA with appropriate hydrophilic solvents; usually ethanol precipitation
- Disadvantages of technique: requires ultracentrifuge, use of mutagenic / toxic ethidium bromide, long centrifugation time (24 - 28 hours)
Ethanol precipitation / salting out
- Uses high aqueous salt concentrations to separate proteins and other cellular contaminants from nucleic acids
- Alcohol is added to aqueous solution and nucleic acids are removed by centrifugation
- DNA is polyanionic due to large number of negatively charged phosphate groups in its phosphodiester linked backbone; dissolves well in highly polar / high dielectric constant solvents like water
- DNA typically does not form ionic bonds with dissolved cations in aqueous solutions, as water molecules form a hydration shell around most ions preventing ionic bond formation and DNA precipitation; an exception is calcium cations, which rapidly bind and precipitate phosphate groups out of aqueous solutions, forming a precipitate that is difficult to resolubilize
- Common cations used in DNA precipitation include Na+, NH4+, Li+
- Ethanol and isopropyl alcohols are less polar than water, having a dielectric constant of 24.3 (ethanol) vs 80.1 for water
- Adding ethanol to an aqueous DNA solution to a concentration of 64% or greater allows the phosphodiester DNA backbone to form ionic bonds with cations, resulting in DNA precipitation
- For optimal DNA precipitation, the concentration of cations must be sufficient to form ionic binds with the DNA but not have so much salt that it precipitates out of solution by itself; conversely, too few cations results in poor DNA precipitation and a low DNA recovery
- Optimal DNA precipitation is achieved at room temperature; storage of precipitation mix at low temperatures lowers precipitation efficiency
- Low precipitation temperatures are often used because they lower nucleic acid degradation from trace nucleases in solution
- In general, shorter DNA fragments require longer periods to precipitate
- DNA recovery is facilitated by higher molecular weight DNA and high DNA concentrations
- DNA precipitation is facilitated by centrifugation, usually at 12,000 g and higher, at 0 - 4 °C
- Resulting DNA pellet is washed, often with 70% ethanol or isopropyl alcohol to remove remaining salts
- Often alcohol precipitation purification is followed by other purification techniques, such as organic extraction or CsCl centrifugation to achieve greater nucleic acid purity
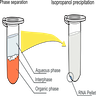
Organic extraction
- Mix crude cell preparations in a denaturing aqueous solution (often guanidinium thiocyanate or 2-mercaptoethanol) with a roughly equal volume of a phenol-chloroform solution and centrifuge the sample
- Isoamyl alcohol may be added to phenol-chloroform solution to reduce foaming during extraction
- Following vigorous mixing, the denatured proteins and lipids partition into the lower organic phase or at the aqueous / organic phase interface, while the nucleic acids partition into the upper aqueous phase allowing separation
- Aqueous sample is often extracted using phenol-chloroform several times for greater nucleic acid purity
- Alternatively, the aqueous solution is washed several times with an isoamyl alcohol-chloroform solution to remove residual phenol
- Nucleic acids are often further purified by ethanol precipitation
- Must use polypropylene tubes, as phenol-chloroform dissolves polystyrene tubes
- Phenol is toxic and can cause severe chemical burns, so protection (gloves, safety glasses, lab coat) and a fume hood are required
- pH of solution must be correct (pH ~7.5), as DNA will partition into organic phase with pH ~5
- Relatively cumbersome and slow compared to other recent organic extraction methods
- Advantage: relatively long nucleic acid polymers can be extracted, useful for some research and molecular diagnostic applications
Silica adsorption
- Uses a silica microchannel matrix with a specific salt concentration and pH (Wikipedia: DNA Separation by Silica Adsorption [Accessed 29 May 2018])
- DNA within a previously prepared cell lysate or other crude nucleic acid mixture preferentially binds to the walls of the microchannels in the presence of a high salt buffer
- DNA binds to the silica under conditions of high ionic strength and a pH at or below the pKa of the silica surface silanol (SiOH) groups
- Multiple washing steps with an alcohol based wash buffer remove other cellular constituents, particularly proteins
- Later the pure DNA is removed with a specific low salt elution buffer; presently, this technique is relatively expensive, limiting its application
- Many automated DNA extraction machines employ a silica matrix for DNA capture and purification
PCR Inhibitors
- PCR inhibitors are commonly encountered; removing these containments is vital to insure good PCR amplification
- Chemical structure of PCR inhibitors and mechanism(s) of inhibition show enormous variation
- Inhibitors include excess salts such as KCl and NaCl, detergents / hydrophobic chemicals (phenol, SDS, chloroform, sodium deocyholate and sarkosyl), inappropriate heating, loss of reagents required for PCR (such as Mg++, Taq polymerase denaturation, etc.), alcohols (ethanol and isopropyl alcohol), heparin, bromophenol blue, agarose, inosine, humic acids, deoxyuridine, excess nucleic acids in the amplification reaction, various polysaccharides (OpenWetWare: PCR Inhibitors [Accessed 29 May 2018])
- Since inhibitors can be either hydrophilic or hydrophobic (or both), DNA purification protocols usually involve subjecting the crude nucleic acid mix to both hydrophilic and hydrophobic washing steps
Commercial DNA extraction machines
- Many DNA extraction machines are commercially available, allowing high throughput simultaneous processing of many samples
- Most commercial extractors use an initial lysis step to free nucleic acids, followed by binding to a solid phase such as magnetic glass beads or a silica membrane
- Solid phase is then extensively washed to remove soluble contaminants and the nucleic acids are removed with heating, high salt buffers or other methods to lower their affinity for the solid matrix
- Separation of the solid phase bound nucleic acids is often facilitated by centrifugation or pull down of magnetic beads in a magnetic field
- Commercial DNA extraction machines were first introduced in the early 1990s and now are vastly more efficient, with higher throughput volumes, better extraction and final purity and lower cost per extraction
Analyzing purity
- Spectrophotometric analysis (A260/280 method)
- Quick and inexpensive
- Theory: nucleic acids absorb significant ultraviolet (UV) light at 260 nm, with the degree of ultraviolet light absorption being directly related to the nucleic acid concentration, by the Beer-Lambert law (Wikipedia: Beer-Lambert Law [Accessed 29 May 2018])
- Extinction coefficient for single stranded DNA is 0.027 (ug/ml)-1 cm-1, for double stranded DNA is 0.020 (ug/ml)-1 cm-1, for single stranded RNA is 0.025 (ug/ml)-1 cm-1
- Based on absorption at 260 nm, an optical density gradient or OD of 1.0 corresponds to 50 ug/ml for double stranded DNA
- Proteins strongly absorb UV light at 280 nm, mainly due to the amino acids tyrosine, tryptophan, cysteine; thus, the concentration of contaminating proteins within a nucleic acid sample can be calculated from the absorbance ratios at 260 and 280 nm (A260/280)
- A260/280: ~1.8 for pure DNA samples; ~2.0 for pure RNA samples
- A260/280 falls with increasing protein contamination, is ~0.57 for pure proteins in solution without nucleic acids
- Nucleic acid samples should be free of phenol, which absorbs strongly at 270 nm
- Pure nucleic acids should have zero absorbance at 330 nm; absorption at 330 nm and above indicates visible light absorption by particulates in solution
- Nucleic acid quantification with fluorescent dyes
- Can quantify nucleic acids by measuring fluorescence intensity of dye bound nucleic acids
- Useful to quantify relatively minute nucleic acid concentrations, helpful if sample has too many absorbing A260 contaminants
- Accuracy due to specificity of nucleic acid binding and fluorescence
- Method: DNA or RNA binding dye is added to a nucleic acid solution; solution is loaded into agarose gel or a surface like a plastic wrap
- Nucleic acid samples of known concentration are added to unknown sample and nucleic acid concentration is estimated by comparing unknown to known samples
- Although somewhat subjective, this method works well for samples with low concentrations of nucleic acids
- This method can be made more exact using microplates or cuvettes measurements against a standard curve on a fluorescent photometer
- Dyes include ethidium bromide, SybrGreen 1, cyanine dyes, RiboGreen, PcoGreen and DAPI
Diagrams / tables
Videos
Cesium chloride density gradient centrifugation